
Note: For larger files, you might experience the five-minute timeout limitation in Lambda. This makes the transformation code portable and allows the Spark jar to be reused if other data sources are added-for example, electronic health records (EHR), billing systems, and other research datasets. The transformation logic is stored in an S3 bucket and is completely de-coupled from the Apache Spark program that runs on EMR and converts the data into OMOP form. This architecture provides a scalable way to use various healthcare sources and convert them to OMOP format, where the only changes needed are in the SQL transformation files. After that message is pushed, it triggers a Lambda function that consumes the message and executes a COPY command from S3 into Amazon Redshift for the appropriate table.As each file is loaded in OMOP form into S3, the Spark program sends a message to an SNS topic that signifies that the load completed successfully.The data is then written to a staging S3 location, where it is ready to be copied into Amazon Redshift.The transformation queries are located in a separate Amazon S3 location, which is read in by Spark and executed on the newly registered tables to convert the data into OMOP form.The files are registered as tables in Spark so that they can be queried by Spark SQL. The MIMIC-III data is read in via an Apache Spark program that is running on Amazon EMR.This includes the Amazon EMR cluster, Amazon SNS topics/subscriptions, an AWS Lambda function and trigger, and AWS Identity and Access Management (IAM) roles. The entire infrastructure is spun up using an AWS CloudFormation template.The data conversion process includes the following steps: The following diagram shows the architecture that is used to convert the MIMIC-III dataset to the OMOP CDM. Solution architecture and loading process However, you don’t need access to the MIMIC-III data to follow along with this post. Getting access to the MIMIC-III dataīefore you can retrieve the MIMIC-III data, you must request access on the PhysioNet website, which is hosted on Amazon S3 as part of the Amazon Web Services (AWS) Public Dataset Program. The code was modified to run in Amazon Redshift. This example demonstrates an architecture that can be used to run SQL-based extract, transform, load (ETL) jobs to map any data source to the OMOP CDM. (See the OHDSI Common Data Model repo in GitHub.) In this scenario, the data is moved to AWS to take advantage of the unbounded scale of Amazon EMR and serverless technologies, and the variety of AWS services that can help make sense of the data in a cost-effective way-including Amazon Machine Learning, Amazon QuickSight, and Amazon Redshift. Observational Health Data Sciences and Informatics (OHDSI) provides the OMOP CDM in a variety of formats, including Apache Impala, Oracle, PostgreSQL, and SQL Server. 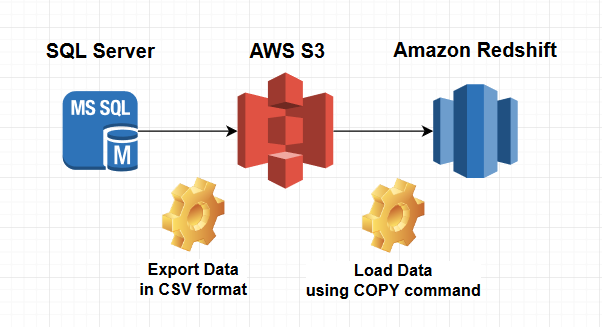
The great advantage of converting data sources into a standard data model like OMOP is that it allows for streamlined, comprehensive analytics and helps remove the variability associated with analyzing health records from different sources. Community resources are available for converting datasets, and there are software tools to help unlock your data after it’s in the OMOP format. The CDM is gaining a lot of traction in the health research community, which is deeply involved in developing and adopting a common data model. The OMOP CDM helps standardize healthcare data and makes it easier to analyze outcomes at a large scale. Note: If you arrived at this page looking for more info on the movie Mimic 3: Sentinel, you might not enjoy this post. This covers the information necessary for processing and storing patient health information (PHI). It describes the architecture and steps for analyzing data across various disconnected sources of health datasets so you can start applying Big Data methods to health research.īefore designing and deploying a healthcare application on AWS, make sure that you read through the AWS HIPAA Compliance whitepaper. This post demonstrates how to convert an openly available dataset called MIMIC-III, which consists of de-identified medical data for about 40,000 patients, into an open source data model known as the Observational Medical Outcomes Partnership (OMOP) Common Data Model (CDM). 
Despite efforts to standardize terminology, some concepts (e.g., blood glucose) are still often depicted in different ways. In the healthcare field, data comes in all shapes and sizes.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
